Note: for strains where we have DNA barcodes we can be reasonably confident of identity, however for those not yet sequenced we rely on morphology
and the original identification, usually made by the depositor. Although CCAP makes every effort to ensure the correct taxonomic identity of strains, we cannot guarantee
that a strain is correctly identified at the species, genus or class levels. On this basis users are responsible for confirming the identity of the strain(s) they receive
from us on arrival before starting experiments.
For strain taxonomy we generally use AlgaeBase for algae and
Adl et al. (2019) for protists.
| Attributes | |
| Authority | (Drezepolski) Nudelman & Triemer 2006 |
| Isolator | Droop (1951) |
| Collection Site | cow's hoof print near Lyme Regis, Dorset, England, UK |
| Climatic Zone | Temperate |
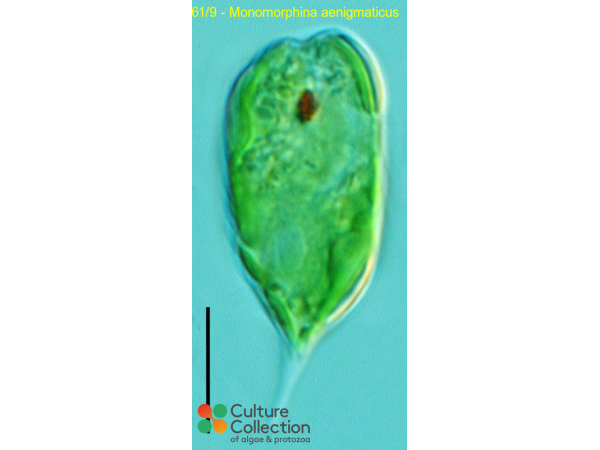
| Notes | Images 1-2 by Bozena Zakrys, Warsaw University and identity amended to P. aenigmaticus May 2001; Renamed as per AlgaeBase after taxonomic revision. |
| Axenicity Status | Bacteria and other organisms present |
| Area | Europe |
| Country | UK |
| Environment | Freshwater |
| GMO | No |
| In Scope of Nagoya Protocol | No |
| ABS Note | Collected pre Nagoya Protocol. No known Nagoya Protocol restrictions for this strain. |
| Collection Date | c 1951 |
| Pathogen | Not pathogenic: Hazard Class 1 |
| Strain Maintenance Sheet | |
| Toxin Producer | Not Toxic / No Data |
| Type Culture | No |
| Taxonomy WoRMS ID | 577248 |
| Equivalent Strains | SAG B1261-9,UTEX 1284 as P. megalopsis |
| Synonyms | Phacus aenigmaticus |
| Formerly Listed in CCAP as | Phacus megalopsis,Phacus aenigmaticus Drezepolski 1921 |
CCAP 1261/9
Monomorphina aenigmaticus
- Product Code: CCAP 1261/9
- Availability: See Availability/Lead Times